2. Getting started
EasyNMR is a web-based solution for handling and processing NMR data with a visual workflow. Here, you can see how easy it is to use EasyNMR!
2.1. Loading data
This example shows how to load NMR data into EasyNMR. Download this dataset (will submit the data in datasets/histcm to Zenodo for public access by users) for solid-state 15N spectrum of L-Histidine Hydrochloride Monohydrate. Alternatively, you can use an NMR dataset of your own. In the L-histidine dataset, you see two data files: histcm.zip which is a zipped Bruker NMR folder, and histcm.csdf which is the same dataset in the CSDM format.
In EasyNMR, you can load NMR data from a variety of formats. Open EasyNMR in your browser by visiting https://easy.csdm.dk. Drag histcm.zip to the white space in the left frame of EasyNMR - which is called Workflow. After dropping the file into EasyNMR, it
loads the zip file,
detects the 1D Bruker spectrum,
creates a CSDM data object from the 1D spectrum, and
attaches a Plot function object to the data object.
Scroll down the Object Contents frame or double click on the Plot object to see the chemical shift spectrum.
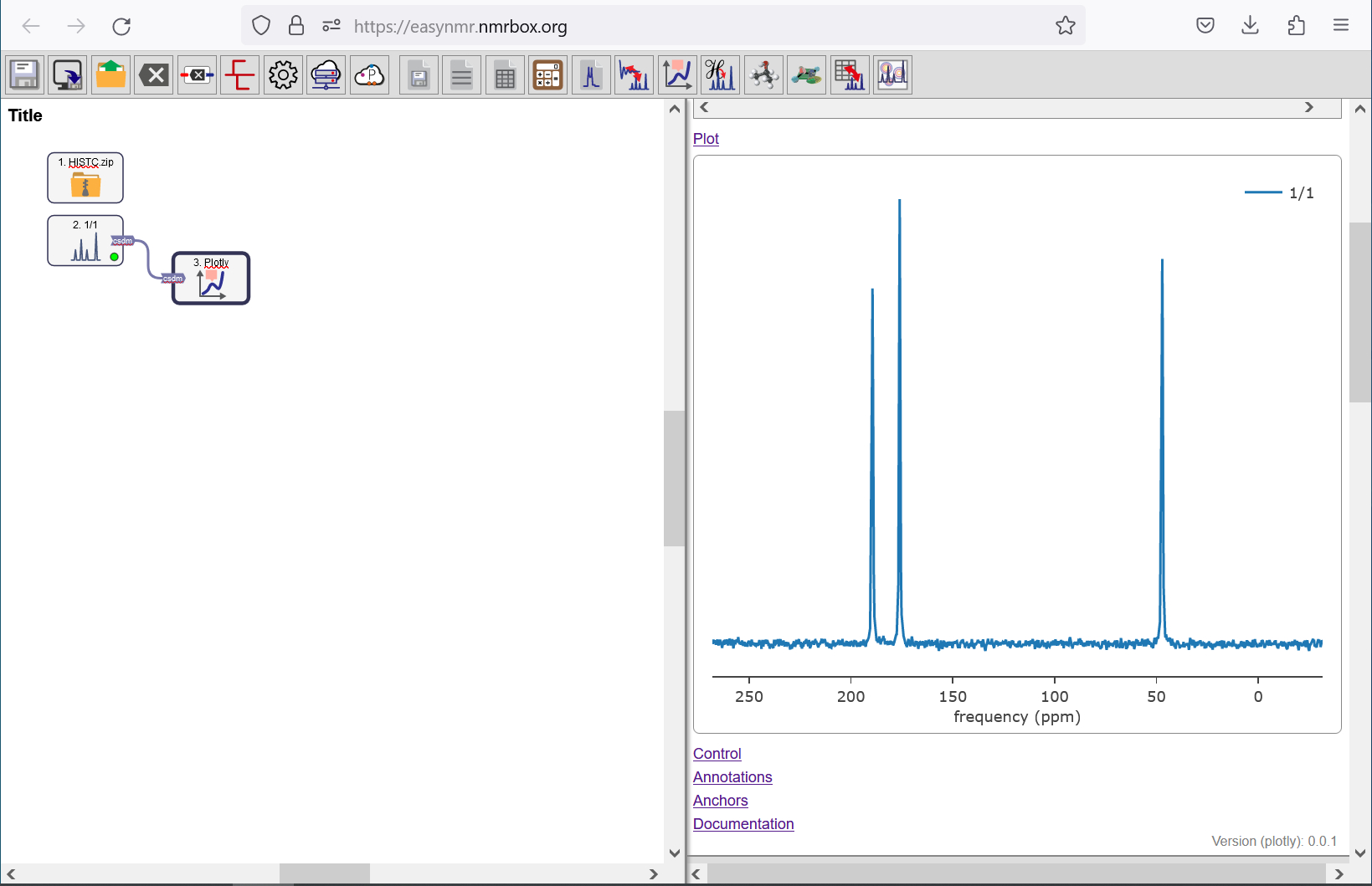
Dragging and dropping HISTC.zip in EasyNMR workflow. Double click on the Plot object to see the spectrum.
Move your cursor over the plot to see the chemical shift corresponding to each peak: \(\delta_{A} = 47 \textrm{ ppm}\) for the amine nitrogen, \(\delta_{\epsilon} = 176 \textrm{ ppm}\) for the nitrogen nuclei farthest from the branch, and \(\delta_{\delta} = 189 \textrm{ ppm}\) for the imidazole nitrogen closest to the branch. You can use Zoom and Pan buttons to explore the spectrum. Click on Autoscale to return to the original view of the spectrum.
2.2. Integration
For integrating peaks, click on the Integrate button of the Plot object to start integration. For each integration, do a single click on the left and then on the right side of the peaks. The relative intensity of each integral will appear beneath each integration bound, with a ratio of [0.95, 1.1, and 1] for [\(\textrm{N}^{A}\), \(\textrm{N}^{\epsilon}\), \(\textrm{N}^{\delta}\)]. Deviation from [1, 1, 1] is due to the cross polarization pulse sequence used in acquiring the spectrum.
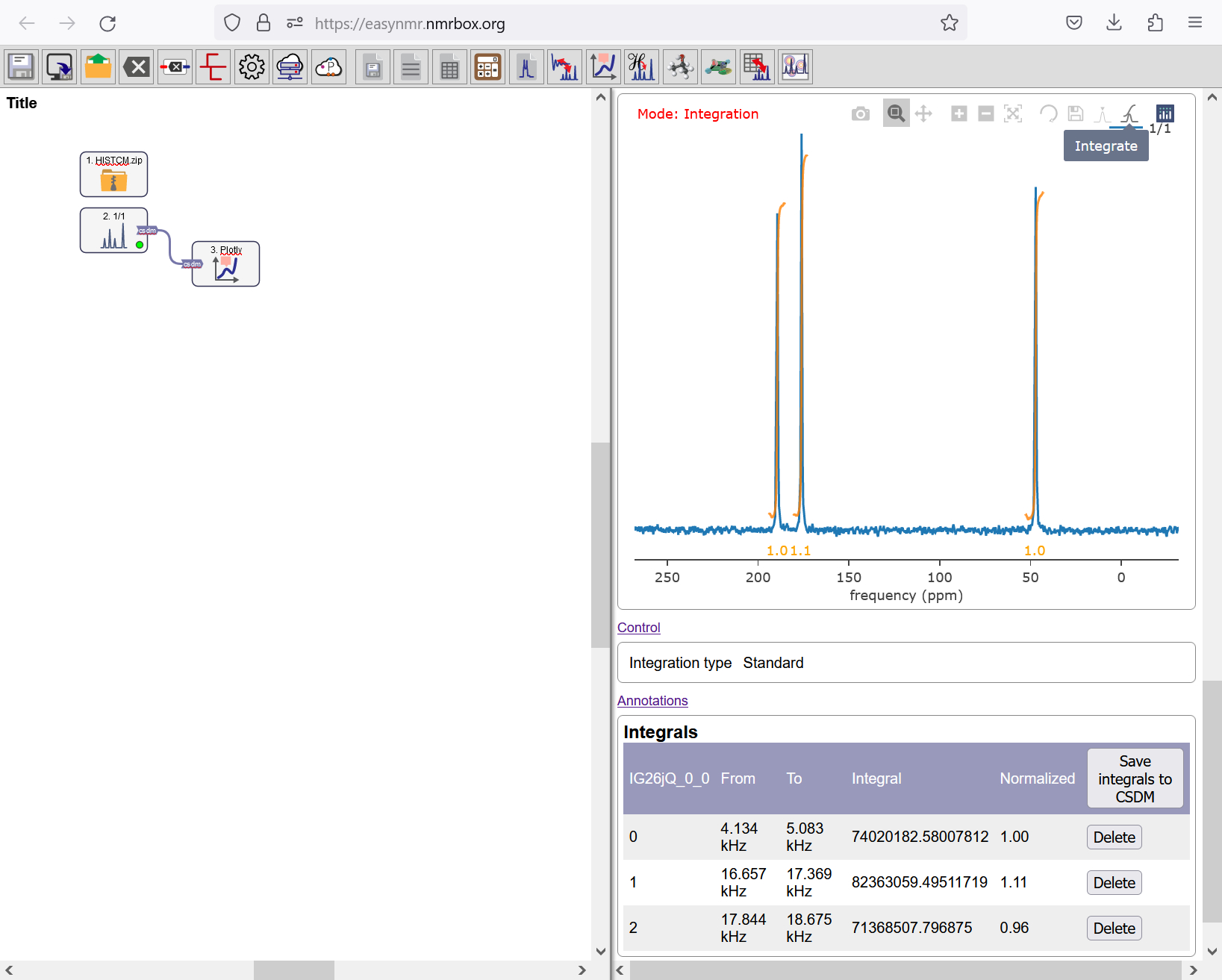
Integration mode with L-histidine hydrochloride monohydrate.
You can see the crystal structure of L-histidine hydrochloride monohydrate by drag and dropping histcm.cif into the Workflow. A CIF Molecule object is created with an attached SIMMOL object. Enable the Ball and Stick option in the Control list. Now you can see the crystal structure of L-histidine hydrochloride monohydrate. Nitrogen nuclei are marked in blue. You can drag the view to rotate the crystal structure.
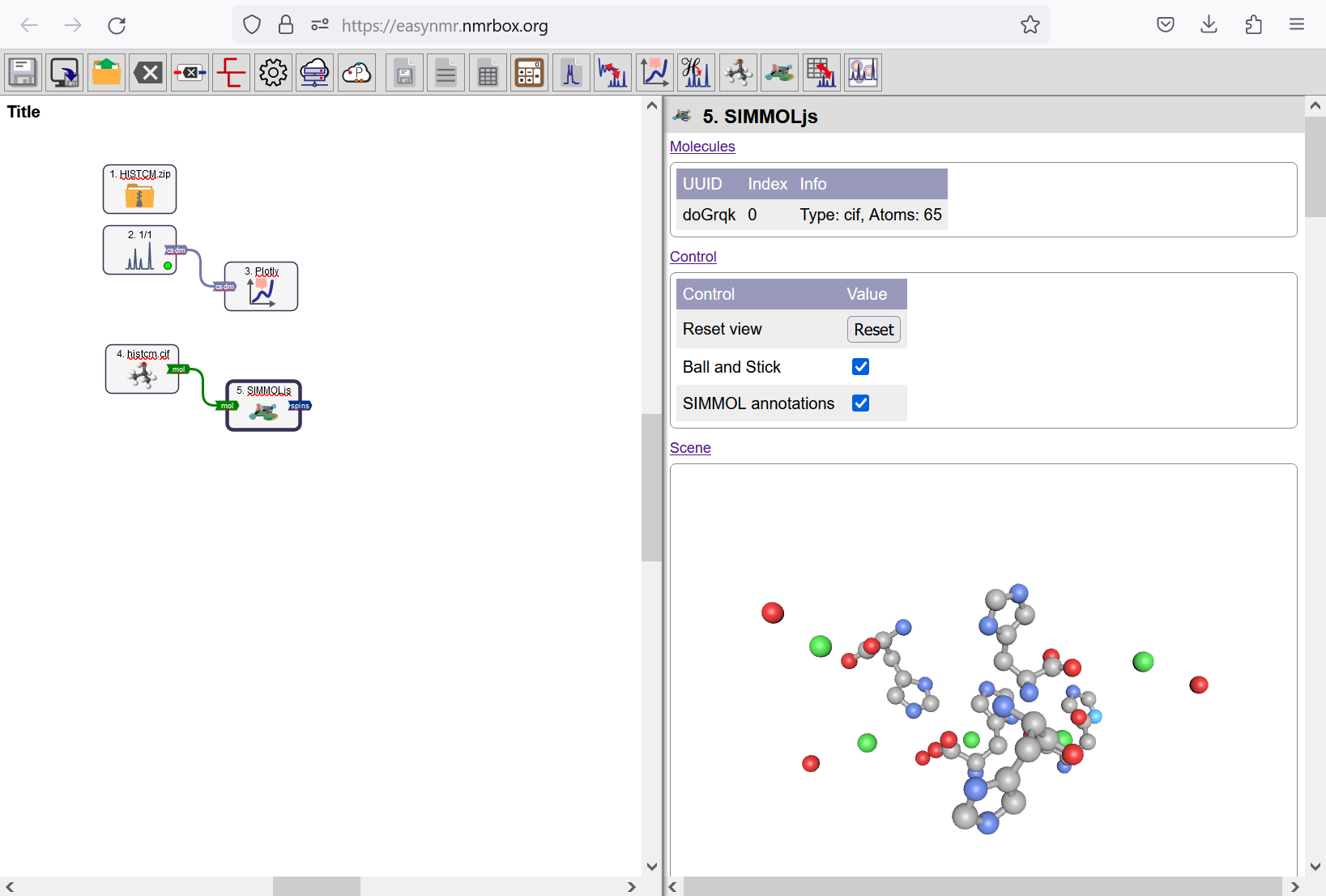
Crystal structure of L-histidine hydrochloride monohydrate shown with the SIMMOL object.
Note
The same NMR data is also stored in histc.csdf in the CSDM file format. EasyNMR internally only stores and processes scientific data in the CSDM format. Dragging and dropping histc.csdf into the Workflow will have the same effect as that of loading the zip dataset.
2.3. Saving and sharing your work
To save your work, or share it with others, simply click on the Export button at the Toolbar. An .easy file will be downloaded by your browser.
Anyone with the file can drag and drop it in an empty EasyNMR window, or load it with the Open button on the Toolbar.
You can share the easy file with your
2.4. Logging in
You do not need to log in for using EasyNMR - with two exceptions:
numerical simulations with the Simulation module - that uses SIMPSON, and
automatic peak picking in the plot object with NMRGlue.
To log in, click on the eServer module. Push the Login button and use your ORCID credentials. If you don’t already have an ORCID account, make it here.
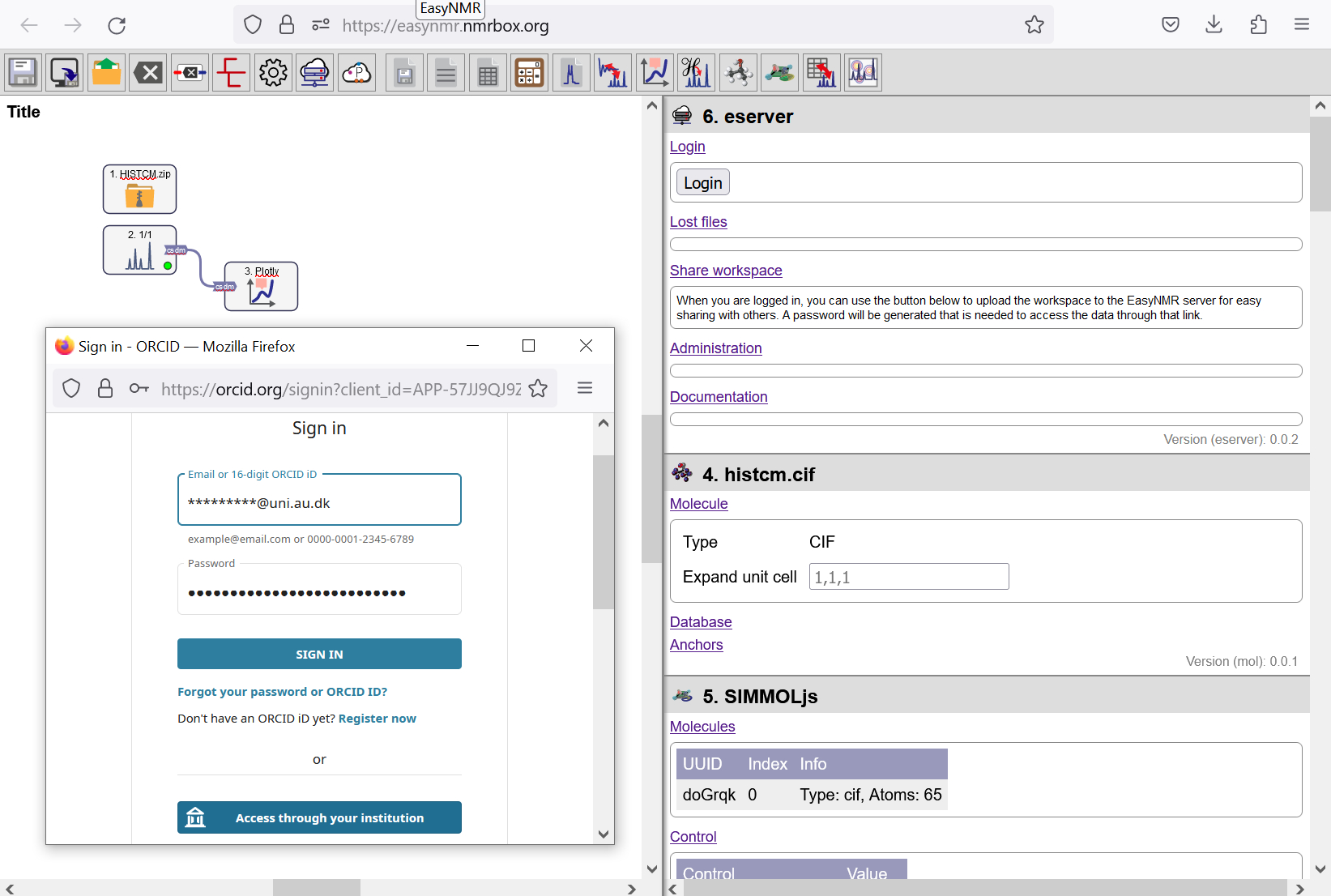
Logging in with eServer and ORCID.
You need to be logged into EasyNMR for some of the tutorials.